|
Learn what it is like to be an undergrad in the Briggs lab!
0 Comments
There are so many postdoctoral fellowship opportunities out there it can be overwhelming at times. Which ones do I apply to? Which ones are the best fit? How many do I apply to? In this blog post, I will not even try to answer these questions, but instead I provide a list of fellowships that I have compiled.
(1) NSF- 3 competitive areas: (1) Biological collections, (2) Broadening participations, (3) plant genomics. Comes with a stipend + research money. (2) La Kretz post-doctoral fellowship- this is associated with the La Kretz center at UCLA. They have only been doing this one for the past 2 or 3 years. Typically, they only take 1-2 new post-docs a year. This has a stipend + some research money. (3) Smith Fellow- This fellowship comes out of the Smithsonian- and is highly geared towards conservation. You need to explicitly say what your conservation plan. About 100 people apply a year, and only 4-5 people get it. Super competitive. If you can get your hands on a the application of a successful fellow- that might increase your chances of getting it because you will have seen the efforts they went to. This seems to have a coherent cohort of fellows and brilliant conservation biologists. (4) Beckman Fellowship- For lifes ciences, including natural and life sciences- but they don't emphasize that. I don't know too much about this one- a friend just mentioned it to me in passing, but you could be located anywhere according to her. This has stipend + research money. (5) Presidential post-doctoral fellowship- This fellowship is shared among the UC schools, Michigan universities, and I think 1 other state. This fellowship strongly emphasizes broadening impacts. This fellowship is particularly competitive- because they typically try to recruit fellows as professors after the postdoctoral fellowship is over. (7) Nature conservancy- Mostly linked to climate change research. I don't know too much about this one. I think the application proposal is super short [1 page]. (8) AAAS fellowship- if you are interested in policy and science- this is the fellowship for you. Competitive and you might have to move to DC. You are essentially paired with an agency or department and learn about the politics and do science. (9) SESYNC postdoctoral fellowship- SESYNC is a synthesis center [similar to NCEAS] in Annapolis, Maryland. (10) NIMBioS postdoctoral fellowship- This fellowship is geared towards merging math and science- and similar to SESYNC and NCEAS- it is a synthesis center (11) USGS Mendenhall research fellowship- This fellowship typically has subcategories- but it doesn't look like they are up yet for this year. I don't think I qualified under any of they categories- so I didn't apply. (12) USGS Powell Center synthesis grant- I've had collaborators apply for this one- I think 25 people apply a year, and they only give out 1 or 2. I don't know too much about this one, but again more math geared. (13) Look on the ecolog listserv. They always advertise for post-doctoral fellowship hirings. They are very specific postings- but might be worth just looking over everyday. (14) Berkeley Miller fellowship. These fellowships are awarded to people who show great promise in their field and who have shortly completed their PhD when they apply. (15) NCEAS postdoctoral fellowship. NCEAS is the world's first science synthesis center and it was launched in 1995. Fellows gain great access to infrastructure and support. (16) Smithsonian tropical research institute fellowships. 3 year postdoc with research money in Panama! (17) Smithsonian museum fellowships. Looks like a lot of opportunities here! (18) Smithsonian environmental research center fellowships. More opportunities! There are likely other postdoctoral fellowships in biology that I have forgotten, but as I come across them- I will keep updating the list. Hope this helps! And good luck! UPDATE: (19) Quantitative postdocs opportunities in the US. This is a compiled spreadsheet by Dr. Allison Barner. (20) L'Oreal postdoctoral fellowship. Note this only provides research funds, not stipend. (21) FIU Distinguished postdoctoral scholar. Along with this fellowship- I highly encourage people to look into the school they are interested at working at to see if there are similar opportunities. (22) National Institute of Health postdoctoral scholarships. (23) Center for Disease Control postdoctoral scholarship. (24) American Museum of Natural History. These fellowships are mostly in evolution and taxonomy, but possible for conservation (25) Liber Ero Fellowships. This is like the Smith Fellowship but only for Canadians. (26) Miller Fellowship at University of California, Berkey (27 LSA Collegiate Fellowship at University of Michigan Recently a colleague asked me, "So I have a twitter account, what do I do now?". Rapidly, I started compiling a list of action items and things to do to get your name out there as a scientist. Here are some thoughts on twitter outreach, networking, and becoming twitter famous (i.e., having lots of followers and people that only know you through twitter):
(1) Follow as many people as you can with similar interests (e.g., biology, stats, disease ecology, etc.), and they will likely follow you back. This will increase the number of followers you have quickly. Just make sure that your little blurb reflects who you are and what you do well- so they can quickly evaluate if they have similar interests. For example, mine says: Post-doc at Michigan State University, Disease Population & Community Ecologist, Herpetologist. Check out my website: http://grazielladirenzo.weebly.com/ Try using lots of key words that other people can identify with- and will say "oh yea! I like population ecology". Personal strategy: The way I look for followers is I go to someone else's twitter page that I know has a lot of followers (for example Karen R. Lips @kwren88), and I just start following her followers or the people that follow her. Go to hashtags or Society journals twitter pages that are interesting to you- and start following people that have similar interested. (2) When you tweet use A LOT of hashtags and use hashtags that people will look at. You are now probably wondering, "How do I know what are the most popular hashtags?". There's a website for that! https://ritetag.com/ Go to that website, and lets say you are going to tweet about science- so put in "science" into the search bar and scroll all the way down to the network map. This map shows you how frequently people visit those hashtags, and give you a sense of "will by tweet get swamped out by all the other people tweeting about science? or will it be visible for a little while?" (3) Plan your tweets ahead of time (check out this blog)- you want the most people as possible to see your tweets and to get your name out there. According to that blog Noon to 1pm is the best time to tweet. I think there is a way to schedule tweets and have them come up at a certain time if you are unavailable. (4) Tweet about websites (that help early career students, advice on how to get a job, mental health problems in academia, how to put together a good presentation, remembering to be color-blind friendly, how to deliver a good seminar, etc.), articles (your newest publications, exciting publications you like, try reading a paper a day for the next year and follow #365papers), and science events you attend (i.e., Euring, ESA, workshops, etc.). Tag people associated with those websites, articles, and events. (5) Not so much a strategy as a fledgling twitterer: Retweeting will help your followers see things that you liked from someone else you follow. This may not necessarily help you gain followers when you are just starting, but it helps spread interesting articles, ideas, websites that you have found to your followers. The value of retweeting increases as you gain followers (I think)- but you can probably dispute that. So take home message: Start following a lot of people (even if you don't know them), start tweeting a lot (websites and articles), use lots of hashtags, and tweet at appropriate times. UPDATE: I had a friend recently ask me, "I have a personal twitter, and I don't really want to give that up. What are the benefits of having a scientific twitter?" Here is my response: (1) When I really started using twitter (i.e., following others)- I swear that I got at least 4 peer-review journals ask me to review papers. I think I got 2 more quickly after that, but I had to decline because I was already doing too much. I don't know if me being more active on twitter was correlated with the increased exposure of my name and my science- but I definitely think it got my name out there. (2) For someone that is shy and has a hard time approaching people at conferences, using twitter has made it is easier for me to talk to people (like, "oh- i really liked your post on X"). (3) From the items I was tweeting about (i.e., being color-blind friendly with your slides, how to give a good presentation, open source science, etc.), I was invited talk to the Mozilla Science Fellows and to the Kuris/Lafferty/Torchin parasitology group at UCSB. (4) Lastly- you get to see cool pictures of what other people are doing, people post papers a lot (some of which I may never have stumbled on), and it is a great way to see exciting science at conferences you can not attend by following their hashtags (i.e., #ESA2017; for the Ecological Society of America meeting in 2017). Again- I think there are a lot of advantages to using twitter as a scientist- but it is ultimately up to you to decide if it worth the time and energy. Check out my guest blog post on my new paper on imperfect pathogen detection that came out in Methods in Ecology & Evolution this year.
Recently, I was invited to write a guest blog post for the Wise Lab and Field Blog about my research. In the post, I describe how I became interested in bayesian hierarchical modeling and the things I am doing now to help others struggling with detection issues..
Hope you enjoy the post! On Dec. 15th 2016, I was an invited speaker at the Mozilla Science Lab Project Call [check them out on twitter @MozillaScience] about Open Science, which is currently being tossed around as a new philosophy in the ecology world. But sometimes it is hard to know what “open science” means.
Here, I’m going to tell you a story about open science. What is it? Why should we do it? Why aren’t people doing it? And what can we do to move towards open science? After that, I will tell a success story in open science, and tell you about what I am doing to promote open science in my own work. I. Open science is:
II. Why should we endorse open science? What are the incentives?
III. I’d like to share with you what I would consider a success story in the Open Science community.
IV. Collectively, as a group, we can move towards open science.
V. Take home message
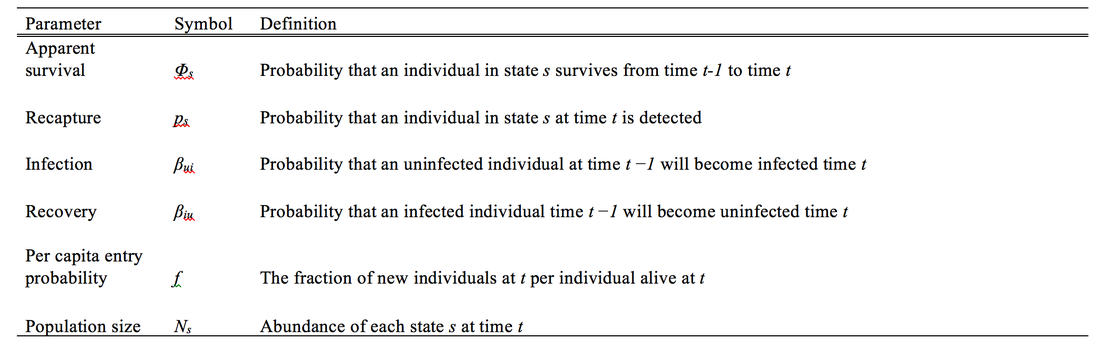
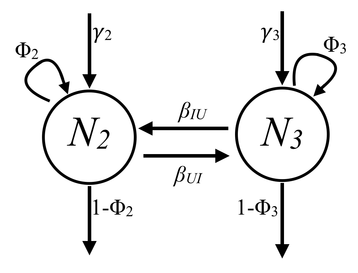
This is going to be a two-part blog post. Here, we will simulate data for a capture-mark-recapture study, and then next time, we will analyze that data in JAGS. Here is the scenario: We are disease ecologists, and we want to understand the disease dynamics of our system. When we capture an individual, we assign it to a certain ‘state.’ A state in our case is a disease status, but can also be defined as a geographic location, an age, a breeding status, etc. Before we begin, I’d like to lay out some information: Our parameters of interest: state-specific apparent survival probability, recruitment probability, transition rates (i.e., movement between two ‘states’), abundance for each state, and population growth rate (depicted in Figure 1). Figure 1. Graphical representation of our model describing the processes contributing to changes in abundance. Parameter names are explained in Table 1. The type of data we ‘collect’: capture-mark-recapture- where each individual is captured, uniquely marked (e.g., elastomer tags, pit tag, toe clips, etc.), and released. Then, each season, new individuals are marked, and old individuals are recaptured. When an individual is captured, we record its state. Our modeling approach: Multi-state Jolly-Seber Model Statistical framework: Bayesian For more information on these models, I highly recommend the book: Bayesian population analysis using WinBUGs by Marc Kéry and Michael Schaub. Table 1 Parameter definitions and symbols. #--------------- R code starts here
#--- Survey conditions n.occasions <- 4 # Number of seasons #---- Population parameters # Super population size of uninfecteds N_U <- 250 # Super population size of infecteds N_I <- 200 # Super population size of all N <- N_U + N_I # Number of states n.states <- 4 # Number of observable states n.obs <- 3 #--- Define parameter values # Uninfected survival probability phi_U <- 0.9 # Infected survival probability phi_I <- 0.7 #------ Entry probability # Entry probability of uninfected hosts gamma_U <- (1/(n.occasions-1))/2 # Entry probability of infected hosts gamma_I <- (1/(n.occasions-1))/2 #------ Transition probability #(Uninfected to Infected) beta_UI <- 0.2 #(Infected to Uninfected) beta_IU <- 0.3 #------ Detection probability # Uninfected detection probability p_U <- 0.9 # Infected detection probability p_I <- 0.7 #---- 1. State process matrix #---- 4 state system # 1. Not entered # 2. Uninfected # 3. Infected # 4. Dead PSI.state <- array(NA, dim = c(n.states, n.states, N, n.occasions-1)) for(i in 1:N){ for(j in 1:n.occasions-1){ PSI.state[,,i,j] <- matrix(c(1 - gamma_U - gamma_I, gamma_U, gamma_I, 0, 0, phi_U * (1- beta_UI), phi_U * beta_UI, 1-phi_U, 0, phi_I * beta_IU, phi_I * (1- beta_IU), 1-phi_I, 0, 0, 0, 1), nrow = 4, ncol = 4, byrow = T) } } # 2. Observational process matrix #----- 3 observed states # 1. Seen uninfected # 2. Seen infected # 3. Not seen PSI.obs <- array(NA, dim = c(n.states, n.obs, N, n.occasions-1)) for(i in 1:N){ for(j in 1:n.occasions-1){ PSI.obs[,,i,j] <- matrix(c( 0, 0, 1, p_U, 0, 1- p_U, 0, p_I, 1- p_I, 0, 0, 1), nrow = 4, ncol = 3, byrow = T) } } #---- Function to simulate capture-recapture data simul.js <- function(PSI.state, PSI.obs, gamma_U, gamma_I, N, N_U, N_I){ n.occasions <- dim(PSI.state)[4] + 1 B_U <- rmultinom(1, N_U, rep(gamma_U, times = n.occasions)) B_I <- rmultinom(1, N_I, rep(gamma_I, times = n.occasions)) # Generate no. of entering hosts per occasion # N is superpopulation size B <- B_U + B_I CH.sur <- CH.p <- matrix(0, ncol = n.occasions, nrow = N) # Define a vector with the occasion of entering the population ent.occ_U <- ent.occ_I <- numeric() for (t in 1:n.occasions){ ent.occ_U <- c(ent.occ_U, rep(t, B_U[t])) ent.occ_I <- c(ent.occ_I, rep(t, B_I[t])) } ent.occ <- c(ent.occ_U, ent.occ_I) # Simulating survival for (i in 1:length(ent.occ_U)){ # For each individual in the superpopulation CH.sur[i, ent.occ_U[i]] <- 2 } for(i in (length(ent.occ_U)+1):length(ent.occ)){ CH.sur[i, ent.occ[i]] <- 3 } for (i in 1:N){ # Probability of capturing the individual during the first occasion if(CH.sur[i, ent.occ[i]] == 2){ CH.p[i, ent.occ[i]] <- 1 * rbinom(1, 1, p_U) }else{ CH.p[i, ent.occ[i]] <- 2 * rbinom(1, 1, p_I) } if (ent.occ[i] == n.occasions) next # If the entry occasion = the last occasion, go next for(t in (ent.occ[i]+1):n.occasions){ # for each occasion from entering 2 last occasion # Determine individual state history sur <- which(rmultinom(1, 1, PSI.state[CH.sur[i, t-1], , i, t-1]) == 1) CH.sur[i, t] <- sur # Determine if the individual is captured event <- which(rmultinom(1, 1, PSI.obs[CH.sur[i, t], , i, t-1]) == 1) CH.p[i, t] <- event } #t } #i #---- Reorder CH.p <- CH.p[ c(which(ent.occ == 1), which(ent.occ == 2), which(ent.occ == 3), which(ent.occ == 4)),] CH.sur <- CH.sur[ c(which(ent.occ == 1), which(ent.occ == 2), which(ent.occ == 3), which(ent.occ == 4)),] # Remove individuals never captured CH <- CH.p * CH.sur cap.sum <- rowSums(CH) never <- which(cap.sum == 0) if(length(never) > 0) { CH.p <- CH.p[-never,] } Nt <- numeric(n.occasions) # Calculate super population size by the simulated data for(i in 1:n.occasions){ Nt[i] <- length(which(CH.sur[,i] > 0)) } #----- CH <- CH.p CH[is.na(CH) == TRUE] <- 0 CH[CH == dim(PSI.state)[1]] <- 0 id <- numeric(0) for(i in 1:dim(CH)[1]){ # For each individual row z <- min(which(CH[i, ] != 0)) # which column is the lowest that does not = 0? ifelse(z == dim(CH)[2], id <- c(id, i), id <- c(id)) # keep a list of individuals only captured on last occasion } CH.sur[CH.sur == 0] <- 1 return(list(CH.p = CH.p[-id, ], CH.sur = CH.sur[-id,], B = B, Nt = Nt)) } # Execute simulation function sim <- simul.js(PSI.state = PSI.state, PSI.obs = PSI.obs, gamma_U = gamma_U, gamma_I = gamma_I, N = N, N_U = N_U, N_I = N_I) #----------------------------# Hope this was helpful and feel free to shoot me an email with any questions at [email protected] As a new feature to my website, I’ve decided to add a Bayesian modeling blog to post code for Bayesian models and make comments on things I wish I knew before I started modeling.
Within the next couple of days, I'll be sharing a project I’ve been part of for several years but just recently started writing the code for. Watch out for my new post on Multi-State Cormack-Jolly-Seber models!! |
Graziella DiRenzoBayesian Analyst sharing my experiences and code Archives
August 2018
Categories |

 RSS Feed
RSS Feed
